Coordination of RNA Polymerase II Pausing and 3′ End Processing Factor Recruitment with Alternative Polyadenylation | Molecular and Cellular Biology

CUT&RUN detects distinct DNA footprints of RNA polymerase II near the transcription start sites | bioRxiv

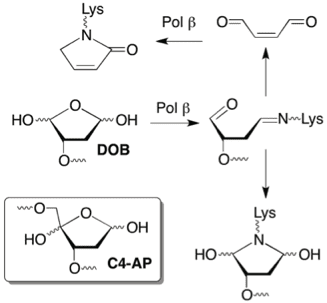
A fidelity mechanism in DNA polymerase lambda promotes error‐free bypass of 8‐oxo‐dG | The EMBO Journal
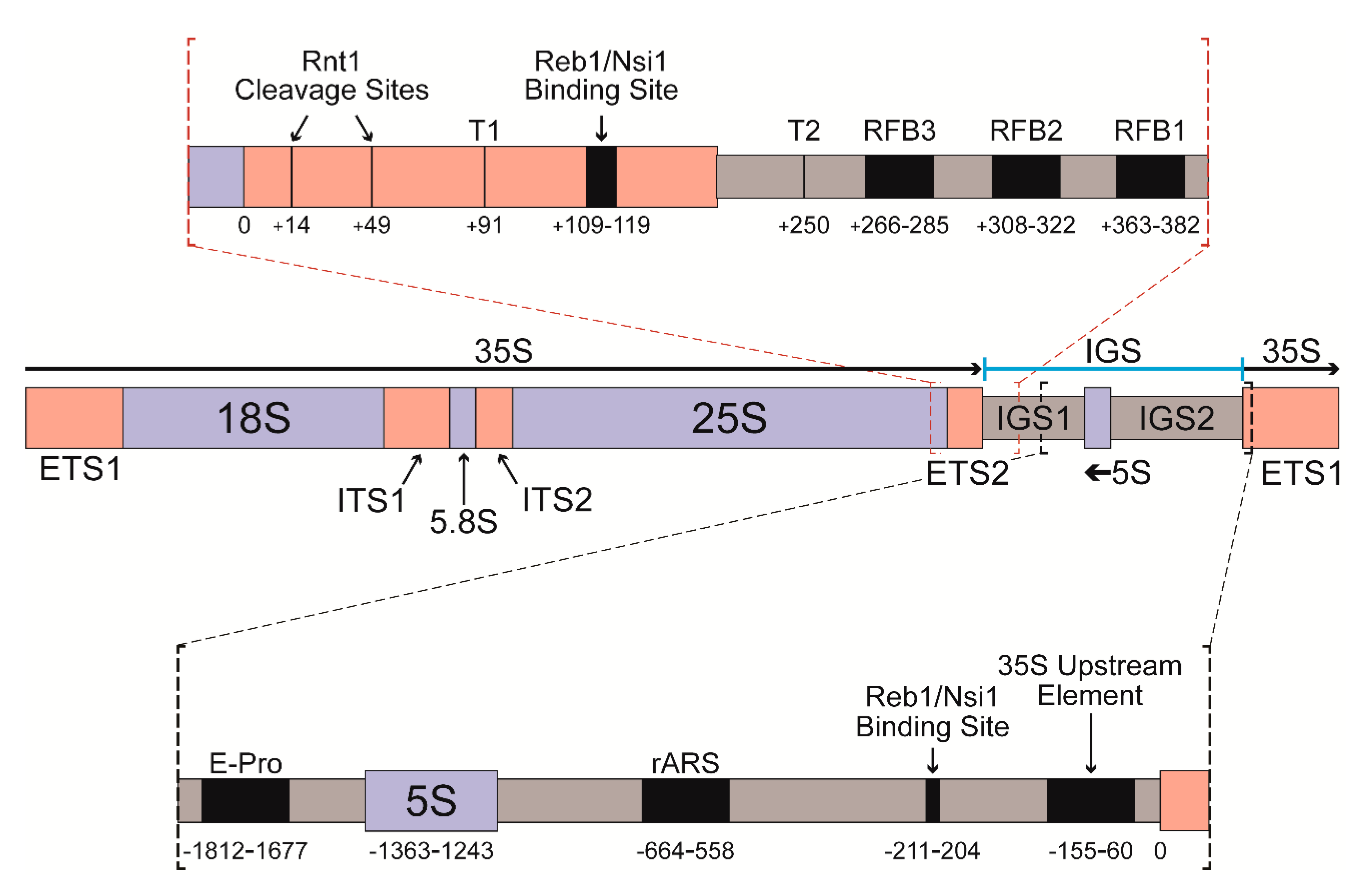
Genes | Free Full-Text | Defining the Influence of the A12.2 Subunit on Transcription Elongation and Termination by RNA Polymerase I In Vivo

Requirement for transient metal ions revealed through computational analysis for DNA polymerase going in reverse | PNAS
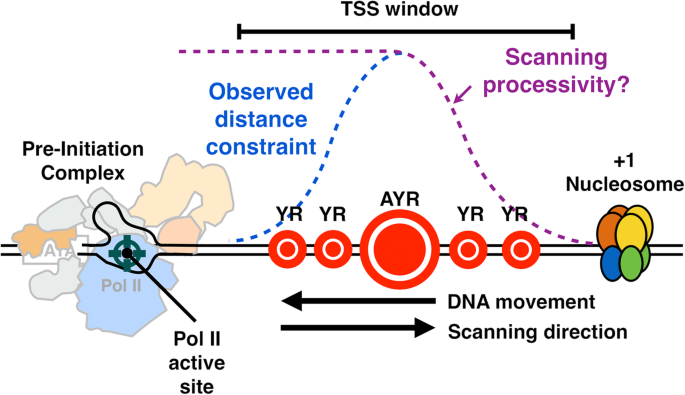
Universal promoter scanning by Pol II during transcription initiation in Saccharomyces cerevisiae | Genome Biology | Full Text
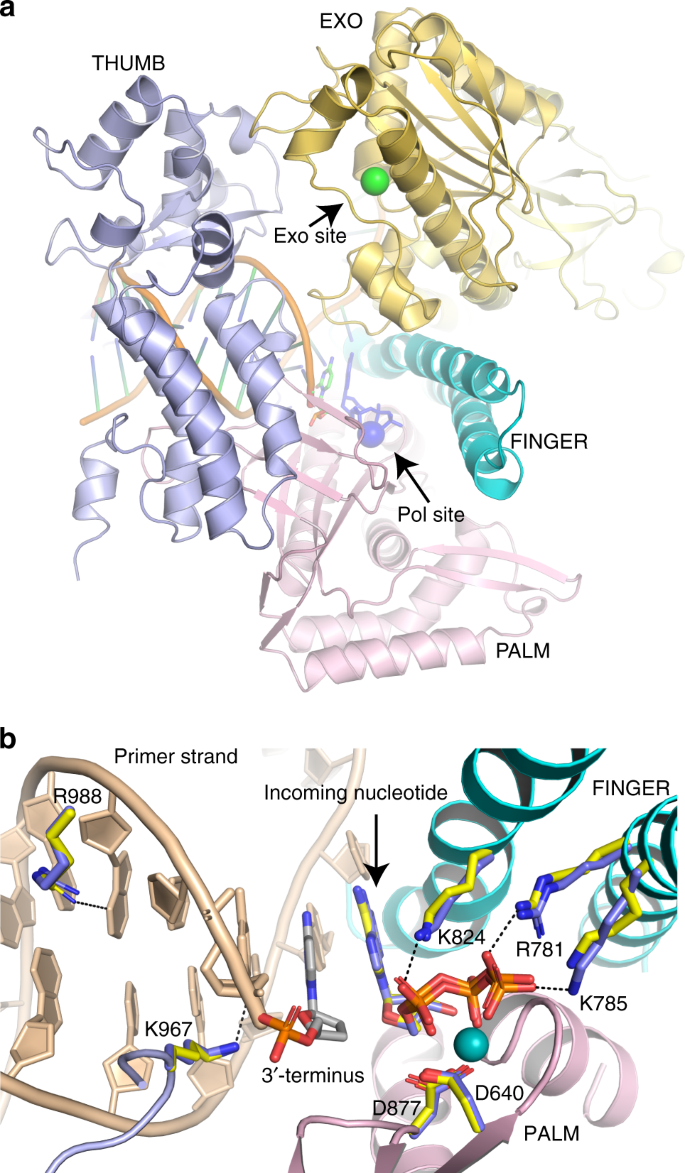
Structural consequence of the most frequently recurring cancer-associated substitution in DNA polymerase ε | Nature Communications

Widespread Backtracking by RNA Pol II Is a Major Effector of Gene Activation, 5′ Pause Release, Termination, and Transcription Elongation Rate - ScienceDirect

Structure and function of the pol II transcription machinery | Taatjes Lab | University of Colorado Boulder

Alternative DNA secondary structure formation affects RNA polymerase II promoter-proximal pausing in human | Genome Biology | Full Text

Promoter-proximal pausing of RNA polymerase II: emerging roles in metazoans | Nature Reviews Genetics

Variations in nuclear localization strategies among pol X family enzymes - Kirby - 2018 - Traffic - Wiley Online Library

POL partners with Army fuels specialists in aircraft refueling initiative > U.S. Air Forces Central > Article Display
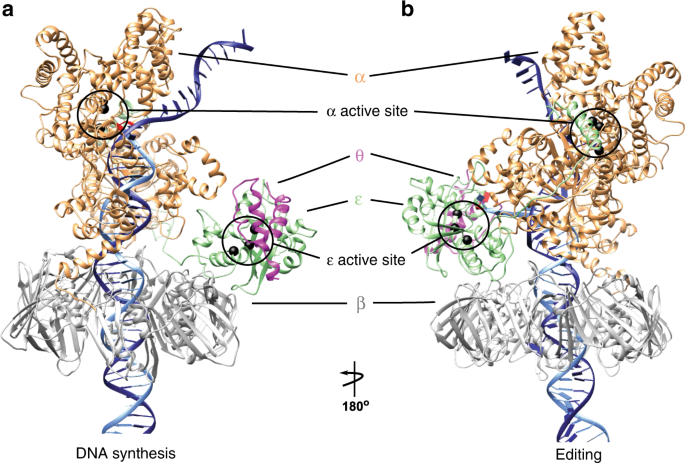
Polymerization and editing modes of a high-fidelity DNA polymerase are linked by a well-defined path | Nature Communications

Probing the structural and molecular basis of nucleotide selectivity by human mitochondrial DNA polymerase γ | PNAS
Mammalian Base Excision Repair: Functional Partnership between PARP-1 and APE1 in AP-Site Repair | PLOS ONE